February 9, 2016
“Clasping” Collaboration
Three CBC Scientists Join Forces to Develop Exceptional-Quality Antibodies Displaying an Unprecedented Mode of Action
Congratulations to three CBC researchers, Shohei Koide and Alex Ruthenburg of The University of Chicago, and Neil Kelleher of Northwestern, who are three senior authors on a collaborative report that was just published in PNAS: “Antigen clasping by two antigen-binding sites of an exceptionally specific antibody for histone methylation.” The paper describes a novel mechanism of a set of exceptionally high-quality antibodies for epigenetics research that the team has developed and meticulously characterized.
Antibodies are essential tools in biomedical research, and the studies of epigenetics, i.e. mechanisms of turning on and off genes, heavily depend on antibodies. Antibodies that selectively capture each of many posttranslational histone modifications (PTMs) of the histone proteins are needed to map these “histone marks” across the genome. However, the scarcity of reliable antibodies has hindered progress in the field. These histone marks, particularly methylation of the amino acid lysine, present a formidable challenge, because an antibody needs to discriminate the minute difference of a single methyl group (CH3–) and distinguish different positions within the histone proteins.
To overcome this “antibody bottleneck” in epigenetics research, a protein-engineering team led by Shohei Koide, Professor of Biochemistry & Molecular Biology at the University of Chicago, generated a set of recombinant antibodies to trimethylated histone H3. The collaborating three CBC groups subjected these antibodies to extensive characterization and identified two of these antibodies that bind to trimethylated histone H3 marks with exquisite specificity. Equally important, unlike animal-derived antibodies commonly used in the field, these recombinant antibodies immediately offer robustness and reproducibility while simultaneously reducing reliance on animals for scientific advancement.
The big surprise came when the antibody-antigen binding was analyzed biochemically and structurally. It has been widely accepted that conventional antibodies are equipped with two copies of the antigen-binding fragment (Fab), each Fab being an independent unit containing a single and complete antigen-recognizing site (Figure A). In other words, the stoichiometry of antigen-Fab binding is typically 1:1 and appears to be evolutionarily conserved. This paradigm has been a guiding principle for developing antibodies as reagents and therapeutics. Unexpectedly, the new antibodies displayed profiles indicative of 2:1 stoichiometry. Crystal structures provided further evidence that in the process of antigen binding, the antibodies actually form head-to-head Fab 1-Fab 2 homodimers, “sandwiching” the antigen in between (Figures B and C).
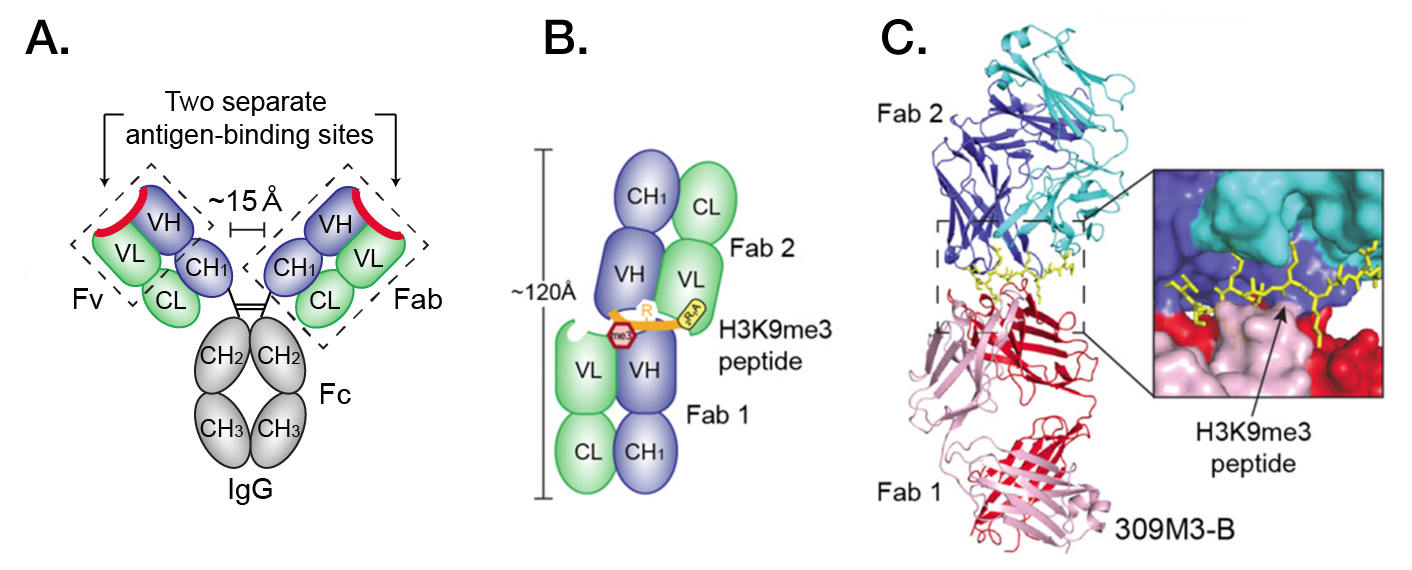
Figure: Classical and “clasping” antigen binding types. (A) Schematic structure of IgG. Two separate binding sites are shown, one per each dimer, each capable of binding one antigen. (B) Schematic structure of one of the generated antibodies (309M3-B) in complex with the trimethylated histone peptide (H3K9me3). Fab 1 and Fab 2 of 309M3-B form an asymmetric head-to-head homodimer “clasping” the H3K9me3 peptide with their individual binding sites. (C) Crystal structure of 309M3-B – H3K9me3 complex. The heavy and light chains in Fab 1 are shown in red and pink, respectively, and those in Fab 2 (the dimerization partner of Fab1) are shown in blue and cyan, respectively. Adapted from Hattori et al., PNAS 2016; published ahead of print February 9, 2016, doi:10.1073/pnas.1522691113. (PNAS Early Edition)
This novel, cooperative antibody-antigen recognition mechanism, where two antigen-binding sites of Fab “clasp” one antigen — hence nicknamed “antigen clasping” — challenges the conventional understanding of antibodies’ mode of action. It also provides incentive and guidance for the development of synthetic antibodies with similarly high performance, because clasping allows the antibodies to use more surfaces to make contact with the antigen and hence discriminate minute details.
The three senior authors on the report (Koide, Alexander Ruthenburg and Neil Kelleher) have strong ties to the CBC. Koide was one of the key participants of the Recombinant Antibody Network supported by the 2012 CBC Lever Award “Center for Production of Affinity Reagents for Human Transcription Factors: Chicago Synthetic Antibody Pipeline,” and has served as CBC Scientific Director for the University of Chicago since 2011. Ruthenburg and Kelleher are both recipients of CBC Recruitment Resources Awards. Ruthenburg joined the University of Chicago in 2010 as a CBC Junior Investigator (joint appointment as Assistant Professor at Molecular Genetics and Cell Biology, and Biochemistry and Molecular Biology Departments). That same year Kelleher was recruited by Northwestern University as a CBC Senior Investigator to become Professor in both the Weinberg College of Arts and Sciences and the Feinberg School of Medicine along with directing the Proteomics Center of Excellence (now with ~25 full-time staff).
“This has been a wonderful collaboration,” said Koide. “It has brought together our truly complementary expertise and the most advanced technologies, which has made it possible to show that these antibodies actually work so well as we claim. The extensive validation is particularly important here, because these antibodies will become gold standard for the field. The clasping mechanism was one of the biggest surprises in my 20-year career studying antibodies. It is fitting that this Chicago-area collaboration has revealed that the architectural principle of Louis Sullivan, ‘Form Follows Function,’ also applies at the molecular level.”
“These antibodies are truly amazing, clearly among the best, if not the best, antibodies that we have used,” added Ruthenburg. “I am excited that, with these revolutionary tools, we are able to discover exciting new epigenetic phenomena and develop diagnostic methods.”



Three CBC collaborators (from the left): Shohei Koide, Neil Kelleher and Alex Ruthenburg.
This work was supported by National Institutes of Health (NIH) Grants R21 DA025725 and RC1 DA028779 (to S.K.) and GM067193 (to N.L.K.). B.D.S. acknowledges funding from the W. M. Keck Foundation. S.K., A.J.R., and N.L.K. acknowledge funding from the Chicago Biomedical Consortium, with support from The Searle Funds at The Chicago Community Trust.
Publication attributed to the CBC funding: Antigen clasping by two antigen-binding sites of an exceptionally specific antibody for histone methylation. Hattori T, Lai D, Dementieva IS, Montaño SP, Kurosawa K, Zheng Y, Akin LR, Świst-Rosowska KM, Grzybowski AT, Koide A, Krajewski K,
Strahl BD, Kelleher NL, Ruthenburg AJ, Koide S. PNAS 2016; published ahead of print February 9, 2016, doi:10.1073/pnas.1522691113.
(PNAS Early Edition)
