July 3, 2019 | Jola Glotzer
HupY in Streptococcus colonization and heme utilization on mucosal surfaces
CBC Postdoctoral Award to Laura Cook and Michael Federle, UIC, leads to the identification of a novel pathway that bacteria utilize to colonize mucosal surfaces
Congratulations to Laura Cook and Michael Federle, UIC, on a recent publication in MBio, titled “Transcriptomic Analysis of Streptococcus pyogenes Colonizing the Vaginal Mucosa Identifies hupY, an MtsR-Regulated Adhesin Involved in Heme Utilization.” The authors acknowledge partial funding of the published work by a CBC Postdoctoral Research Award that the team received in 2014, and was granted a renewal for another year in 2015, titled: “RNASeq Analysis of the Transcriptome of Streptococci Colonizing the Vaginal Tract.”
Using genome-wide transcriptome sequencing (RNASeq), the authors show that two genes — hupYZ — are highly expressed in the human pathogen Streptococcus pyogenes during vaginal colonization. Further analysis shows that HupY, previously characterized as an adhesin in Streptococcus, binds heme and affects intracellular iron concentrations. As HupY protein has no known heme binding domains, the authors propose it is a novel heme binding protein, important to bacterial iron homeostasis and mucosal colonization.
A former postdoc in Federle lab, Laura Cook is currently an Assistant Professor at SUNY-Binghamton, NY. Michael Federle, Associate Professor in the Department of Medicinal Chemistry and Pharmacognosy, UIC, has many other ties to the CBC including being a recipient of two CBC HTS Awards in 2015 and 2013 and a Catalyst Award in 2013. Michael is also a current member of the CBC Catalyst Review Board (see below). The CBC congratulates all authors of the current study and thanks Michael for his many efforts in supporting the CBC.
Publication attributed to CBC funding*:
Cook LCC, Chatterjee N, Li Y, Andrade J, Federle MJ, Eichenbaum Z. Transcriptomic Analysis of Streptococcus pyogenes Colonizing the Vaginal Mucosa Identifies hupY, an MtsR-Regulated Adhesin Involved in Heme Utilization. MBio. 2019 Jun 25;10(3). (PubMed)
ABSTRACT
Streptococcus pyogenes (group A streptococcus [GAS]) is a serious human pathogen with the ability to colonize mucosal surfaces such as the nasopharynx and vaginal tract, often leading to infections such as pharyngitis and vulvovaginitis. We present genome-wide transcriptome sequencing (RNASeq) data showing the transcriptomic changes GAS undergoes during vaginal colonization. These data reveal that the regulon controlled by MtsR, a master metal regulator, is activated during vaginal colonization. This regulon includes two genes highly expressed during vaginal colonization, hupYZ. Here we show that HupY binds heme in vitro, affects intracellular concentrations of iron, and is essential for proper growth of GAS using hemoglobin or serum as the sole iron source. HupY is also important for murine vaginal colonization of both GAS and the related vaginal colonizer and pathogen Streptococcus agalactiae (group B streptococcus [GBS]). These data provide essential information on the link between metal regulation and mucosal colonization in both GAS and GBS.
Importance
Colonization of the host requires the ability to adapt to an environment that is often low in essential nutrients such as iron. Here we present data showing that the transcriptome of the important human pathogen Streptococcus pyogenes shows extensive remodeling during in vivo growth, resulting in, among many other differentially expressed genes and pathways, a significant increase in genes involved in acquiring iron from host heme. Data show that HupY, previously characterized as an adhesin in both S. pyogenes and the related pathogen Streptococcus agalactiae, binds heme and affects intracellular iron concentrations. HupY, a protein with no known heme binding domains, represents a novel heme binding protein playing an important role in bacterial iron homeostasis as well as vaginal colonization.
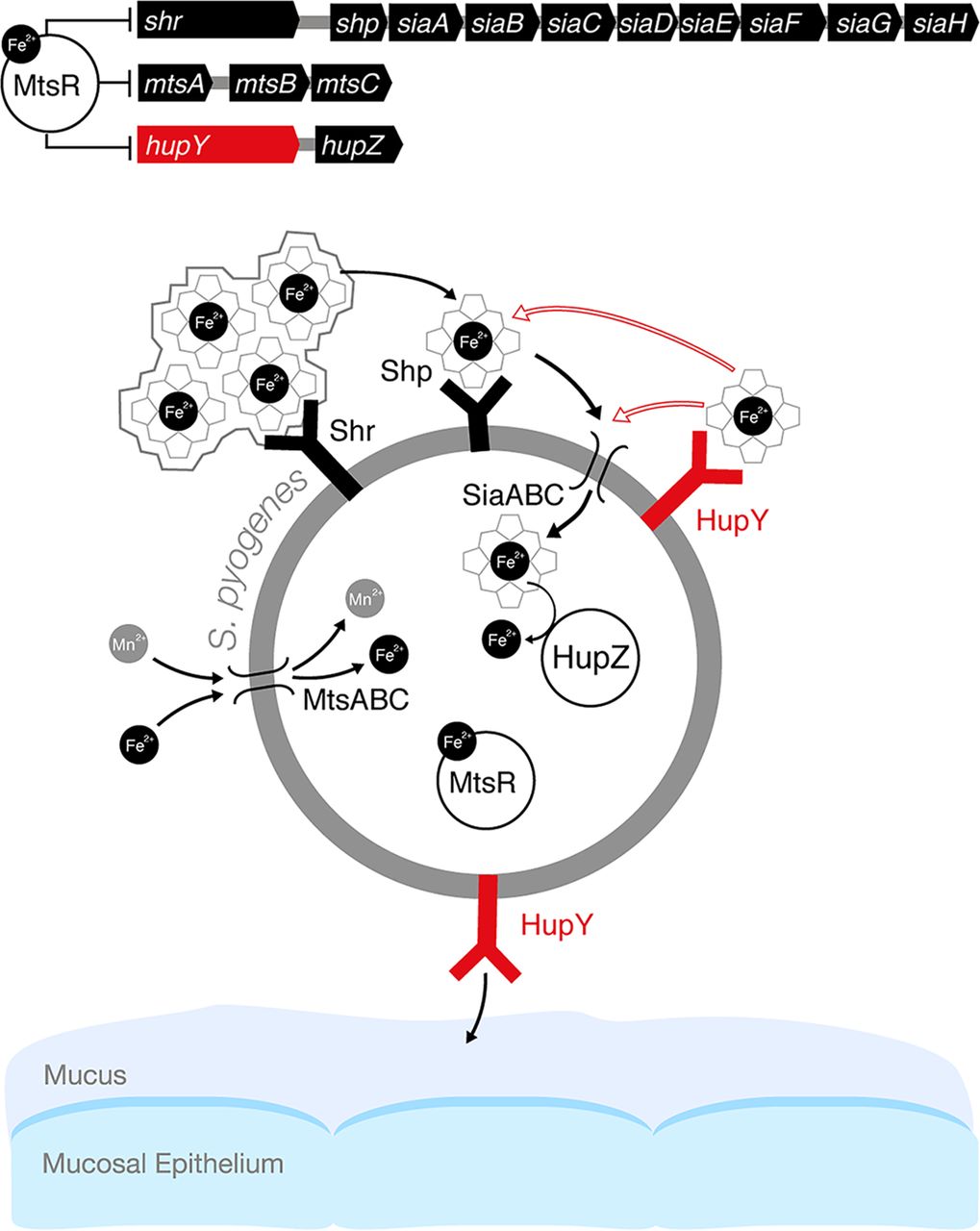
Model of the role of HupY in GAS colonization and heme utilization on mucosal surfaces. In the presence of high levels of iron, iron-bound MtsR represses a large number of genes, including the sia operon, which includes the genes for the siaABC heme importer as well as shr, and shp heme binding proteins. In addition, MtsR represses the manganese and iron mtsABC transporter, hupZ, and hupY. MtsR also downregulates the mtsABC gene cluster in a manganese-dependent manner (not shown in the model). Under iron-depleted conditions such as on mucosal surfaces, MtsR repression is relieved, leading to upregulation of these operons. Shr captures heme (from host hemoproteins or the environment) and delivers it to Shp and subsequently to SiaABC for import into the cell. Once inside the cell, iron can be liberated from heme by the HupZ enzyme. HupY, coregulated with HupZ, is a surface protein that has the ability to bind heme. We propose that HupY binds heme to allow transport to HupZ, either through the Shp/SiaABC heme import pathway or through another mechanism. Open arrows indicate possible heme transfer pathways. HupY also plays a role in mucosal colonization, possibly as an adhesin important for binding to host cells.(Source: mbio.asm.org)
ACKNOWLEDGMENTS
RNA sequencing was performed by the University of Chicago Genomics Facility, and analysis was performed by the Bioinformatics Core Facility at the Center for Research Informatics, University of Chicago. We acknowledge Gabriel Morton-Cook for help with model figure design.
This work was supported in part by American Heart Association Greater Southeast Affiliate Grant-in-Aid 15GRNT25600006 (Z.E.), NIH grant AI091779 and the Burroughs Wellcome Fund Investigators in Pathogenesis of Infectious Diseases (M.J.F.), NIH F32AI110047-01 (L.C.C.C.), and Binghamton University Structural Startup Funds (L.C.C.C.). Portions of this project were supported by a Chicago Biomedical Consortium postdoctoral research grant (L.C.C.C.).
Featured CBC Community member(s):
Laura Cook and Michael Federle, UIC
- 13th Annual CBC Symposium (2015):
▸ The Unseen Majority: Microbes in Health and Disease
Michael Federle (UIC) – Invited speaker - HTS Award (2015):
▸ Reversing Antibiotic Resistance with Compounds that Potentiate Antimicrobial Drugs
PIs: Michael Federle and Tiara Pérez Morales, (UIC) - *CBC Postdoctoral Research Award (2014, renewed in 2015):
▸ RNASeq Analysis of the Transcriptome of Streptococci Colonizing the Vaginal Tract
▸ Renewal
PIs: Laura Cook and Michael Federle (UIC) - CBC Postdoctoral Research Award Review (ad-hoc; 2014-2016):
Michael Federle (UIC) – Reviewer - Catalyst Award (2013):
▸ Immunotherapy-Mediated Interference of Bacterial Quorum Sensing
PIs: Michael J. Federle (UIC) and Matthew Tirrell (UChicago) - HTS Awards (2013):
▸ Signal Jamming of Bacterial Communication: an HTS approach
PI: Michael Federle (UIC) - CBC Scholar Award (2012-2013):
▸ Meet the Scholar Chaitanya Aggarwal
Michael Federle (UIC) – Mentor - CBC Scholars “Loop Connections” Seminar (2012):
▸ CRISPR Loci and Signal Interference in Bacteria
Michael Federle (UIC) – Invited speaker
CBC Catalyst Review Board (CRB; 2016-1017; 2018-present):
▸ Current Membership
Michael Federle (UIC) – Board Member
ARTICLES PUBLISHED IN THE PAST ABOUT THE FEATURED CBC COMMUNITY MEMBER(S):
February 14, 2018
▸ Tricking bacteria to remain non-hostile may become a method of choice in preventive medicine—the Federle lab at UIC shows how
November 5, 2017
▸ Three UIC researchers and CBC awardees featured in the UIC College of Pharmacy Annual Report 2017

