August 1, 2019 | Jola Glotzer
Protein purification “decision tree”
CBC Senior Investigator Neil Kelleher, NU, contributes to a review in Nature Methods on best practices for protein analysis by top-down mass spectrometry
Congratulations to Neil Kelleher, NU, for contributing to a recent review published in Nature Methods, “Best practices and benchmarks for intact protein analysis for top-down mass spectrometry.” Written collaboratively with other members of the Consortium for Top-Down Proteomics, the article provides a “decision tree” (see below) — a comprehensive set of guidelines to purification and analysis of native proteins.
Kelleher is the Walter and Mary Elizabeth Glass Professor of Chemistry, Molecular Biosciences, and Medicine at NU. He is also recognized as a CBC Senior Investigator due to his recruitment to NU in 2010 with help from the CBC Recruitment Resources Funds Award. Kelleher’s other CBC awards and community contributions are listed below the article. The CBC is proud to have played a role in Kelleher’s recruitment and to have helped support his research over the years.
Publication linked to the CBC funding*:
Donnelly DP, Rawlins CM, DeHart CJ, Fornelli L, Schachner LF, Lin Z, Lippens JL, Aluri KC, Sarin R, Chen B, Lantz C, Jung W, Johnson KR, Koller A, Wolff JJ, Campuzano IDG, Auclair JR, Ivanov AR, Whitelegge JP, Paša-Tolić L, Chamot-Rooke J, Danis PO, Smith LM, Tsybin YO, Loo JA, Ge Y, Kelleher NL, Agar JN. Best practices and benchmarks for intact protein analysis for top-down mass spectrometry. Review. Nat Methods. 2019 Jul;16(7):587-594. (PubMed)
ABSTRACT:
One gene can give rise to many functionally distinct proteoforms, each of which has a characteristic molecular mass. Top-down mass spectrometry enables the analysis of intact proteins and proteoforms. Here members of the Consortium for Top-Down Proteomics provide a decision tree that guides researchers to robust protocols for mass analysis of intact proteins (antibodies, membrane proteins and others) from mixtures of varying complexity. We also present cross-platform analytical benchmarks using a protein standard sample, to allow users to gauge their proficiency.
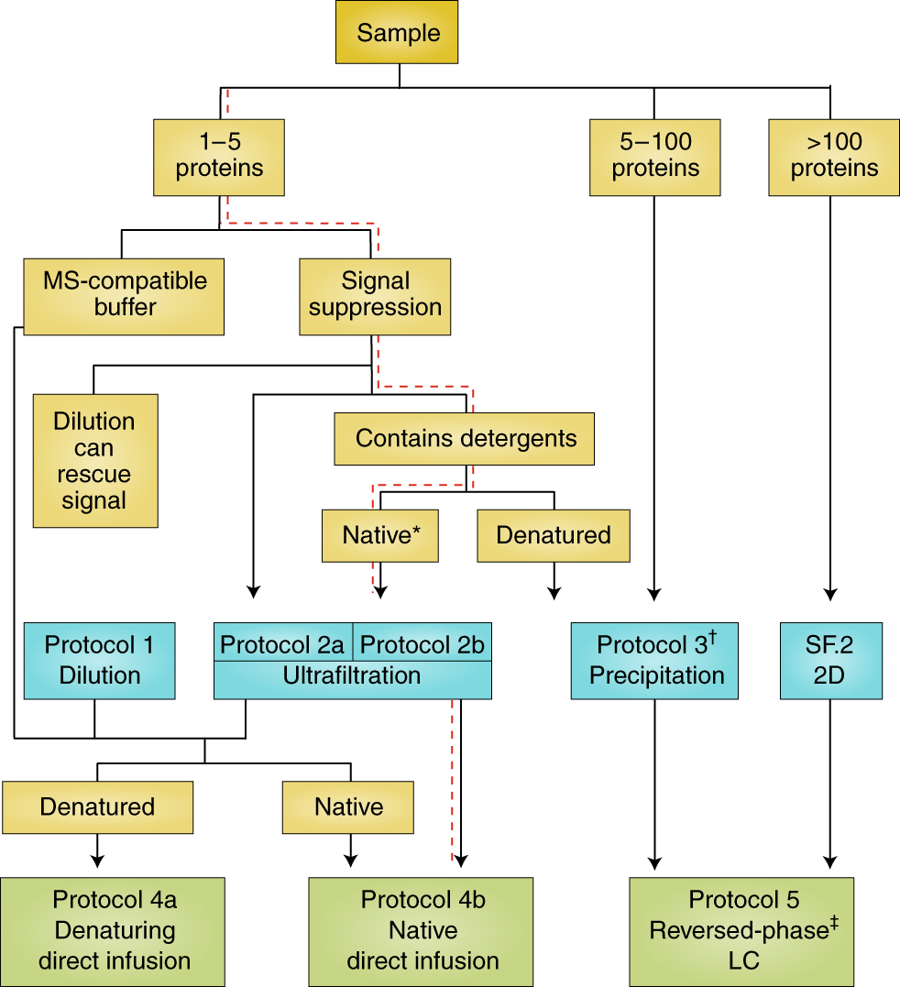
Decision tree for intact protein sample clean-up, preparation and analysis. (Source: www.nature.com)
Featured CBC Community member(s):
Neil Kelleher, NU
- CBC Exploratory Workshop (2013):
▸ The CBC Exploratory Workshop on Cellular Heterogeneity
Neil Kelleher (NU) – Workshop Organizer and Speaker - CBC Science Day (2011):
Neil Kelleher (NU) – Science Day Speaker - CBC Catalyst Award (2010):
▸ Phosphoproteomic Analysis of NADPH Oxidase Activation
PIs: Neil Kelleher (NU) and Richard Ye (UIC) - *CBC Recruitment Resources Award (2010):
▸ CBC Senior Investigator
Neil Kelleher (NU) - CBC Seminar (2010):
▸ Top Down Mass Spectrometry: Has Its Decade Now Come?
Neil Kelleher (NU, UIUC then) – Seminar Speaker
ARTICLES PUBLISHED IN THE PAST ABOUT THE FEATURED CBC COMMUNITY MEMBER(S):
July 8, 2019
▸ Defining the NSD2 interactome
Two CBC Awardees, Jonathan Licht and Neil Kelleher, identify posttranslational modification crosstalk that may play a role in carcinogenesis
July 8, 2019
▸ New insight into liver cancer treatment
CBC Awardee Rick Silverman and a CBC Senior Investigator, Neil Kelleher, NU, collaborate on developing therapeutics for hepatocellular carcinoma (HCC)
June 17, 2019
▸ Copper centers revealed with top-down mass spectrometry
Three CBC Awards contribute to today’s publication in Nature Communications!
May 13, 2019
▸ Battling epigenetic lymphomas
CBC Senior Investigator Neil Kelleher, NU, contributes to a new publication identifying a potential novel therapeutic pathway to treat the so called EZH2 dysregulated lymphomas
May 7, 2019
▸ Proteoforms explained
A recent review in Proteomics, co-authored by a CBC Senior Investigator and proteomics expert, Neil Kelleher, NU
August 9, 2018
▸ Top-down Proteomics
CBC Senior Investigator, Neil Kelleher, NU, explains the advantages of “top-down” versus “bottom-up” proteomics in early cancer detection and progression
November 13, 2017
▸ New insights into regulation of gene expression: work from the Shilatifard lab (NU) with contributions from two CBC scientists, Neil Kelleher and Jeffrey Savas.
October 5, 2017
▸ CBC Senior Investigator, Neil Kelleher, NU, deciphers molecular assembly of a gut toxin, colibactin
February 9, 2016
▸ “Clasping” Collaboration
Three CBC Scientists Join Forces to Develop Exceptional-Quality Antibodies Displaying an Unprecedented Mode of Action

