October 8, 2019 | Jola Glotzer
sADPL profiling of proteome
In a fresh off the press PNAS paper, CBC awardee Ray Moellering, UChicago, describes a novel technique called soluble activity-dependent proximity ligation (aADPL) that may significantly improve proteome analysis.

Raymond Moellering, UChicago, senior author on the PNAS publication, acknowledges the CBC support of his published research
Congratulations to the Moellering team at UChicago on a new publication, “Ultrasensitive, multiplexed chemoproteomic profiling with soluble activity-dependent proximity ligation,” published yesterday (Oct. 7) in PNAS.
The team reports on the development of a novel technique called soluble activity-dependent proximity ligation or sADPL which, quickly and with ultrasensitive specificity, profiles whole protein families based on their enzymatic activity in complex protein samples obtained from cells, fluids, and tissues. When combined with quantitative PCR, aADPL profiling can detect active proteins at picogram levels in unfractionated proteome.
Specifically, sADPL is shown to be very effective in rapid quantifying protein–drug interactions in blood samples and in measuring protein activity in patient tumor samples — literally on a bench and in a matter of hours. Overall, the authors make a compelling proof-of-concept demonstration of the power of the aADPL profiling and the applicability of the method to versatile whole proteome analyses of biological and clinical samples.
The senior author on the paper, Raymond Moellering, PhD, is Assistant Professor at the Department of Chemistry at UChicago. Moellering has several ties to the CBC (see below). He attributes the current study to the CBC funding he received in 2017 via a CBC Catalyst Award. Co-author Ernst Lengyel, MD, PhD, UChicago, is also affiliated with the CBC. Lengyel is Professor and Chairman of the Department of Obstetrics and Gynecology, section Gynecologic Oncology, and co-director of the Ovarian Cancer Research Laboratory at UChicago. Lengyel’s CBC Awards and other ties to the CBC are listed below.
Publication attributed to *CBC funding:
Li G, Eckert MA, Chang JW, Montgomery JE, Chryplewicz A, Lengyel E, *Moellering RE. Ultrasensitive, multiplexed chemoproteomic profiling with soluble activity-dependent proximity ligation. PNAS. First published October 7, 2019. [Epub ahead of print] (doi.org)
ABSTRACT:
Chemoproteomic methods can report directly on endogenous, active enzyme populations, which can differ greatly from measures of transcripts or protein abundance alone. Detection and quantification of family-wide probe engagement generally requires LC-MS/MS or gel-based detection methods, which suffer from low resolution, significant input proteome requirements, laborious sample preparation, and expensive equipment. Therefore, methods that can capitalize on the broad target profiling capacity of family-wide chemical probes but that enable specific, rapid, and ultrasensitive quantitation of protein activity in native samples would be useful for basic, translational, and clinical proteomic applications. Here we develop and apply a method that we call soluble activity-dependent proximity ligation (sADPL), which harnesses family-wide chemical probes to convert active enzyme levels into amplifiable barcoded oligonucleotide signals. We demonstrate that sADPL coupled to quantitative PCR signal detection enables multiplexed “writing” and “reading” of active enzyme levels across multiple protein families directly at picogram levels of whole, unfractionated proteome. sADPL profiling in a competitive format allows for highly sensitive detection of drug–protein interaction profiling, which allows for direct quantitative measurements of in vitro and in vivo on- and off-target drug engagement. Finally, we demonstrate that comparative sADPL profiling can be applied for high-throughput molecular phenotyping of primary human tumor samples, leading to the discovery of new connections between metabolic and proteolytic enzyme activity in specific tumor compartments and patient outcomes. We expect that this modular and multiplexed chemoproteomic platform will be a general approach for drug target engagement, as well as comparative enzyme activity profiling for basic and clinical applications.
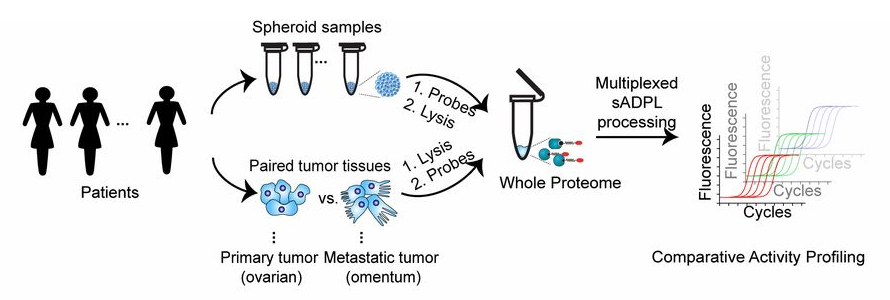
Multiplexed sADPL activity profiling in patient-derived tumor samples. Schematic depiction of sADPL profiling of cancer spheroids and primary tumor and metastatic tumor tissues. (Source: PNAS)
ACKNOWLEDGMENTS:
We thank all the patients who kindly donated samples. We thank C. He for access to instrumentation, P. Dauer for assistance with mouse PBMC isolation, and S. Ahmadiantehrani for proofreading assistance. This work was supported by NIH Chemical Biology Interface Training Grant 2T32GM008720-16 ( to J.E.M.), the Marsha Rivkin Foundation (M.A.E.), National Cancer Institute Grant R01 CA111882 (to E.L.), NIH Grants R00 CA175399 and DP2 GM128199-01 (to R.E.M.), the Chicago Biomedical Consortium supported by Searle Family Funds (R.E.M.), University of Chicago Cancer Center Support Grant P30 CA014599, and the Duchossois Family Institute at the University of Chicago.
Featured CBC Community member(s):
Raymond Moellering, UChicago
- CBC Accelerator Review Board (2018-present):
Raymond Moellering (UChicago) — Board Member - 15th Annual CBC Symposium (2017):
▸ Small Molecule Discovery
Raymond Moellering (UChicago) — Symposium Speaker - *CBC Catalyst Award (2017):
PIs: Raymond Moellering (UChicago) and Sarki Abdulkadir (NU) for the project:
▸ Defining Prostate Tumor Malignancy with Activity-Dependent Pet Probes
Ernst Lengyel, UChicago
- CBC Postdoctoral Research Award (2014; renewal, 2015):
▸ Differentiating between benign and malignant ovarian tumors using targeted contrast agents for magnetic resonance imaging
PIs: Yilin Zhang (postdoc) and Ernst Lengyel, UChicago - CBC HTS Award (2013):
▸ HTS Using an Organotypic Culture of Ovarian Cancer Metastasis
PIs: Hilary Kenny and Ernst Lengyel, UChicago - CBC Exploratory Workshop (2013):
▸ The CBC Ovarian Cancer Workshop
PI: Ernst Lengyel, UChicago — Workshop Organizer and Speaker
ARTICLES PUBLISHED IN THE PAST ABOUT THE FEATURED CBC COMMUNITY MEMBER(S):
October 7, 2019
▸ Fighting metastatic ovarian cancer
CBC High-Throughput Screening (HTS) Award helps Hilary Kenny and Ernst Lengyel, UChicago, identify three promising ovarian cancer metastasis inhibitors
May 21, 2019
▸ Cells surrounding the cancer promote its spread
Two CBC Awardees, Ernst Lengyel and Raymond Moellering, UChicago, contribute to recent research published in Nature about the role of stroma in facilitating metastasis of ovarian cancer cells
May 4, 2019
▸ Consider slimming down!
CBC affiliate Ernst Lengyel, UChicago, comments in U.S.News on the rise of obesity-related cancer
February 19, 2019
▸ CT45, or cancer/testis antigen 45
Past CBC Awardee, Ernst Lengyel, UChicago, contributes to the discovery of a biomarker that can help predict response to therapy in patient suffering from ovarian cancer
September 28, 2018
▸ Glycogen fuels ovarian cancer spread
Three CBC awardees, Ernst Lengyel, Hilary Kenny and Yilin Zhang, UChicago, co-author new publication in Cell Metabolism
October 6, 2017
▸ Chicago-area holds the fourth highest number of this year’s NIH Director’s New Innovator Awards nation-wide
Three of the four Chicago-area scientists awarded the 2017 NIH Director’s New Innovator Award, Jingyi Fei, Jaehyuk Choi and Raymond Moellering are CBC community members
February 18, 2015
▸ Modeling a 3D Human Ovarian Cancer in vitro to Screen for Metastasis Inhibitors
